CGS offers a wide range of bioinformatic solutions for the analysis of your data that can be tailored to your exact needs whether the data is generated by us or another service provider. From our automated pipelines to building bespoke analysis, we will make sure you can get the most information out of your data. At the planning stage, our experienced bioinformatians can review your design and validate it or advise on possible modifications in order to have the best possible outcome for your experiment. Our team has built up considerable experience in microarray and sequencing data analysis and our automated pipelines were designed using state of the art software and analysis packages. All our automated pipelines generate an analysis report that details and explain each steps of the analysis.
We offer a complete analysis solution from raw data to result and can also help generate quality graphs for your publications.
Below is a brief description of our analysis portfolio. However, if you require specific analysis not listed here please contact us and we can work with you on developing a bespoke analysis pipeline tailored to your requirements.
Applications
Sequencing:
- RNA-Seq: differential gene expression, splice variation and detection of de novo molecules such as long non-coding RNAs.
- Small RNA-Seq: differential gene expression, modification and prediction of microRNAs and small non-coding RNAs.
- DNA-Seq: Chip-seq, amplicon, targeted methylation, exomes, whole-genomes.
- Long Read Sequencing: We use the Oxford Nanopore system for DNA, cDNA and direct RNA sequencing of reads of long reads >50kb.
- Microbiome: We have extensive experience in 16/18S analysis of microbial communities.
- Custom: We can work on a custom sequencing strategy for your particular project.
Arrays:
- Gene Expression (mRNA, miRNA)
- Genotyping
- CNV/LOH
- Methylation
- Custom
Other Services
- Experimental Design Advice
- Automated Pipeline
- Bespoke Pipeline
- Bespoke data visualisation and data analysis.
Example Data
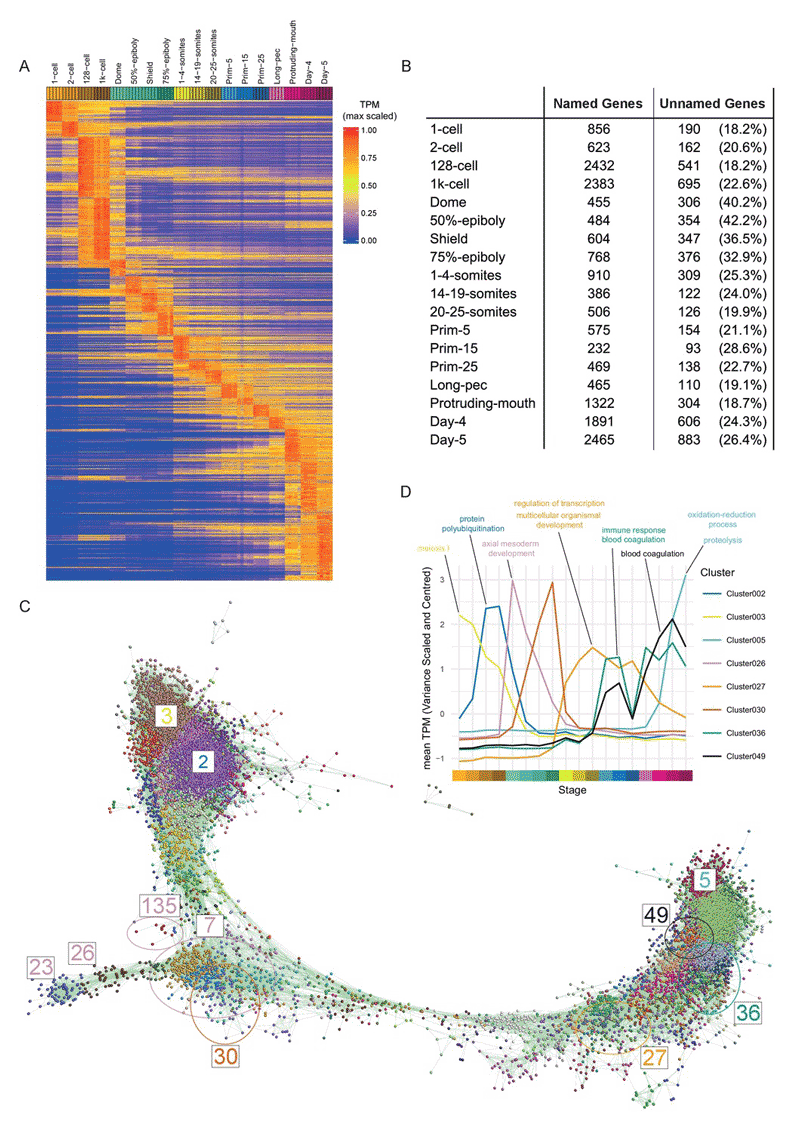
Large-scale mRNA-Seq in early Zebrafish development, including expression mode correlation analysis and graph based clustering of gene-expression profiles using Markov Clustering.